* Welcome *

You are number...
- click the words for navigation -


| F1 Karen (Dim Sum Dolly) | M1 Lucas (EMO boy) |
| F2 Delia (Chua Hong Hong) | M2 Alvin (Guest-Of-Honor) |
| F3 Miki (pied piper) | M3 Zhao Zhi (Mr. BOO) |
| F4 Flavia (Ms. pokey) | M4 Mr. Tung Zhi Kang (Kun-jun virus) |
| F5 Siti (sweet lil thing) | |
| F6 Sajini (she smells nice) | |
| F7 Faith (she gives us hope) |



 |
 |
 |
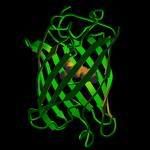
The protein that is responsible for the fluorescent glow in the mouse is called Green Fluorescent Protein, also known as GFP. in this experiment, we will be scaling up colonies of E.coli that is been inserted with GFP.
What is GFP?
GFP is a protein that has a unique β-barrel structure that consists of 11 β-strands + 1 α-helix + chromophore running through the middle. GFP is one of the most commonly used bioreporter of a gene-expression or gene product at the molecular level.
Though GFP was first isolated and purified in the 1960s by Osamu Shimomura from Aequorea victoria, a type of jellyfish., the true potential of GFP was not comprehended until in 1987 when Douglas Prasher first thought of the idea of using GFP as a ‘tag’ that reports whenever a particular protein was being produced by a cell. Since a protein molecule was extremely small to be observed unless under an electron microscope, why not attach a marker so that it can be easily detected even by the naked eye?
So HOW GFP is inserted into the nucleotide sequence and expressed, for example in the case of this experiment, an E.coli??
A simplify diagram of a nucleotide sequence
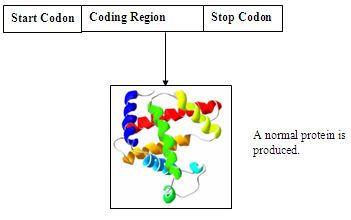
A simplify diagram of a GFP inserted nucleotide sequence
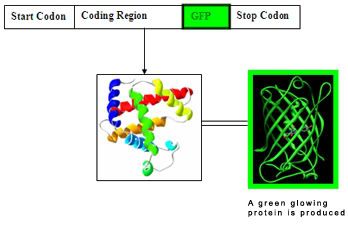
Overall Objectives
The main objective of the whole practical is … *drum roll*… To be able to get an A+++ for our report!!!!!! Hahaha just kidding, well that’s about 80% of the whole objective. The other 20% is to learn and experience the idea of scaling up the growth of E.coli inserted with GFP in a step-wise manner from a seed culture all the way to a bioreactor. At the end of the experiment, we also aim to be able to:
we harvested 10ml of the fermentation broth from a sterile, disposable tube and placed under UV light to check for presence of desired protein produced by the cells, after which we were all shipped to see the history plot and managed to derive some conclusions.
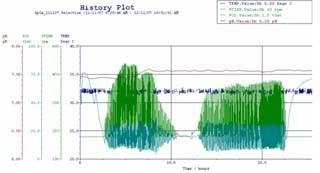
the controlling parameters for the history plot are:
the pH has to be maintained at pH 7.5, since the pH of an actively growing culture will not remain constant for long. pH measurement was done via pH probe which is connected to a pH meter. when the pH of media drop below 7.5, the media becomes acidic, and the computer control system sends signals to pump in sodium hydroxide into fermentor, if pH of media rises above 7.5, causing the medium to be alkaline, sulphuric acid is pumped.
temperature has great influence in the biological processes in a cell through kinetics effects (rate of reaction) and catalytic effects (enzymatic activity). a temperature probe is used to indicate if there are any changes in the temperature of the medium. at 32 oC, desired product GFP folds and fluorescence optimally. any higher temp would denature the protein, the rate of flow of cooling water or steam through the jackets ensures that the temp of fermentor is constant.
E.coli being a living organism, it requires ample amount of air to survive, or the cells will die. hence its an aerobic fermentation. therefore dissolved oxygen here is crucial. the partial pressure of O2 is set to be 20% like in the open environment. when pO2 is not 20%, the rate of injection of air bubbles by the sparger is adjusted.
♥ flavia & faith.